Abstract
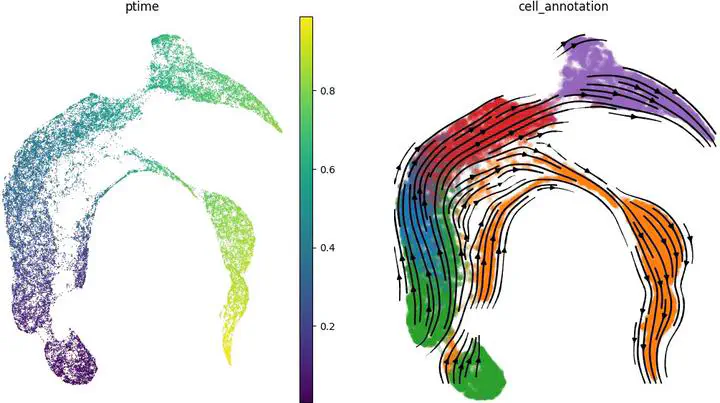
RNA velocity-based methods estimate cellular dynamics and cell developmental trajectories based on spliced and unspliced RNA counts. Although numerous methods have been proposed, RNA velocity-based models vary greatly in their biophysical assumptions, architectures, and use cases. In this work, we introduce a new architecture, CellFlows, which incorporates self-supervised neural dimensionality reduction with the flexibility of neural-based latent time estimation into a mechanistic model, improving model interpretability and accuracy. CellFlows models splicing dynamics to infer gene and context-specific kinetic rates at single-cell resolution and correctly identifies both linear and branching cellular differentiation pathways originating from mouse embryonic stem cells.